Genopathomic profiling identifies signatures for immunotherapy response of lung adenocarcinoma via confounder-aware representation learning
iScience, 2022.
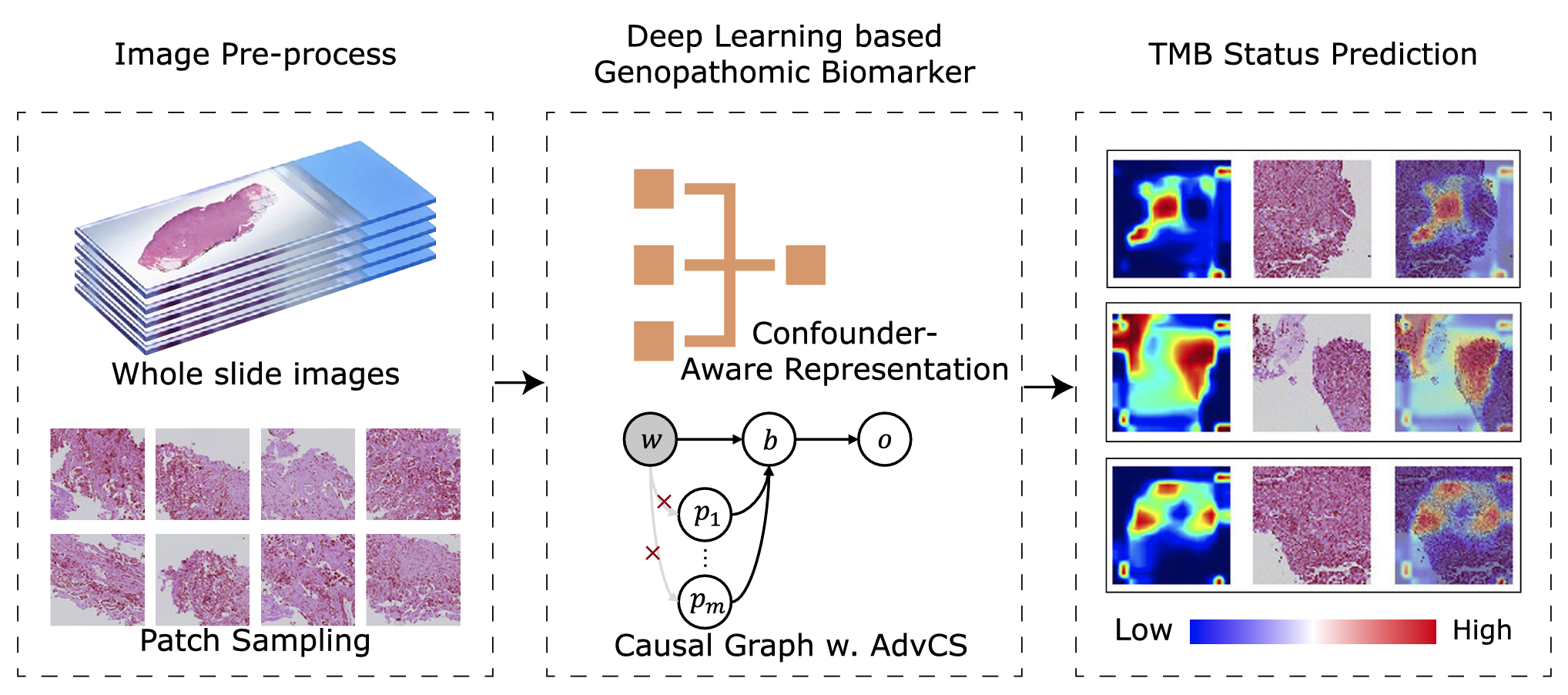
Summary Immunotherapy shows durable response but only in a subset of patients, and test for predictive biomarkers requires procedures in addition to routine workflow. We proposed a confounder-aware representation learning-based system, genopathomic biomarker for immunotherapy response (PITER), that uses only diagnosis-acquired hematoxylin-eosin (H&E)-stained pathological slides by leveraging histopathological and genetic characteristics to identify candidates for immunotherapy. PITER was generated and tested with three datasets containing 1944 slides of 1239 patients. PITER was found to be a useful biomarker to identify patients of lung adenocarcinoma with both favorable progression-free and overall survival in the immunotherapy cohort (p < 0.05). PITER was significantly associated with pathways involved in active cell division and a more immune activating microenvironment, which indicated the biological basis in identifying patients with favorable outcome of immunotherapy. Thus, PITER may be a potential biomarker to identify patients of lung adenocarcinoma with a good response to immunotherapy, and potentially provide precise treatment.
Acknowledgements
The study was supported by the National Natural Science Foundation of China (91959126, 8210071009), the Science and Technology Commission of Shanghai Municipality (20XD1403000, 21Y11913400, 21YF1438200), and the Clinical Research Foundation of Shanghai Hospital Development Center (SHDC2020CR3047B), Clinical Research Foundation of Shanghai Pulmonary Hospital (No. FK1937).