Developmental Stage Classification of Embryos Using Two-Stream Neural Network with Linear-Chain Conditional Random Field
International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI), 2021.
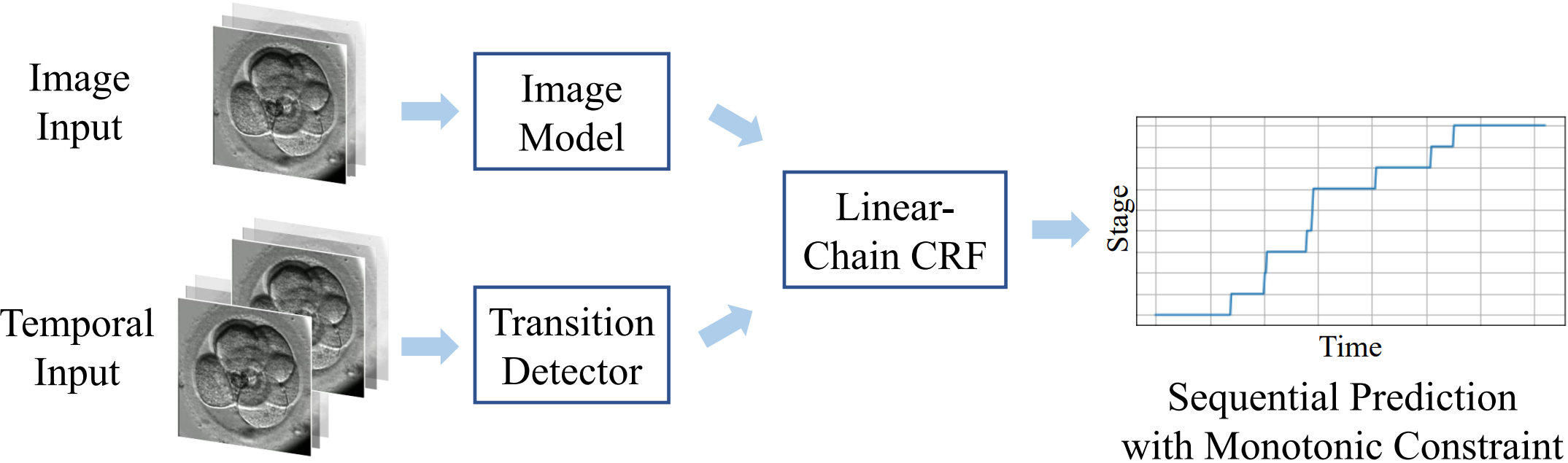
The developmental process of embryos follows a monotonic order. An embryo can progressively cleave from one cell to multiple cells and finally transform to morula and blastocyst. For time-lapse videos of embryos, most existing developmental stage classification methods conduct per-frame predictions using an image frame at each time step. However, classification using only images suffers from overlapping between cells and imbalance between stages. Temporal information can be valuable in addressing this problem by capturing movements between neighboring frames. In this work, we propose a two-stream model for developmental stage classification. Unlike previous methods, our two-stream model accepts both temporal and image information. We develop a linear-chain conditional random field (CRF) on top of neural network features extracted from the temporal and image streams to make use of both modalities. The linear-chain CRF formulation enables tractable training of global sequential models over multiple frames while also making it possible to inject monotonic development order constraints into the learning process explicitly. We demonstrate our algorithm on two time-lapse embryo video datasets: i) mouse and ii) human embryo datasets. Our method achieves 98.1% and 80.6% for mouse and human embryo stage classification, respectively. Our approach will enable more profound clinical and biological studies and suggests a new direction for developmental stage classification by utilizing temporal information.