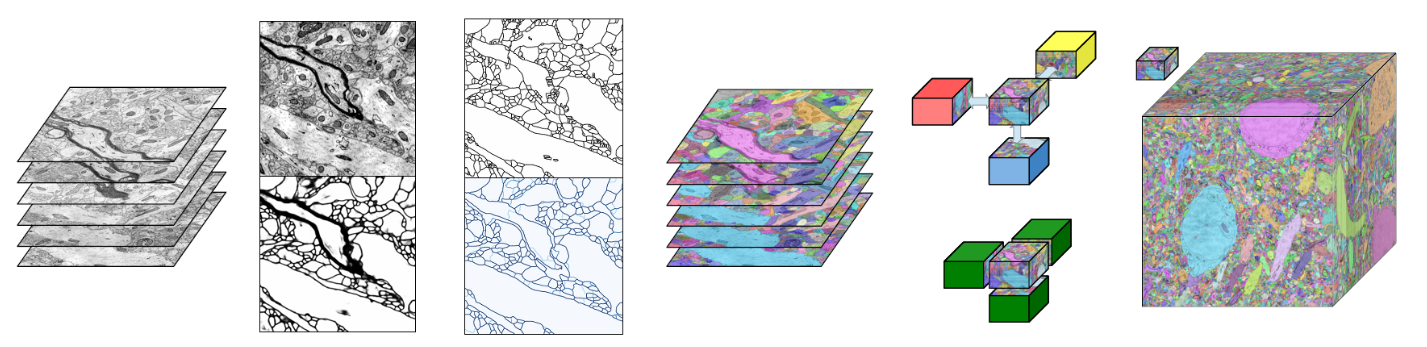
RhoanaNet Pipeline: Dense Automatic Neural Annotation
arXiv preprint arXiv:1611.06973, 2016.
Reconstructing a synaptic wiring diagram, or connectome, from electron microscopy (EM) images of brain tissue currently requires many hours of manual annotation or proofreading. The desire to reconstruct ever larger and more complex networks has pushed the collection of ever larger EM datasets. A cubic millimeter of raw imaging data would take up 1 PB of storage and present an annotation project that would be impractical without relying heavily on automatic segmentation methods. The RhoanaNet image processing pipeline was developed to automatically segment large volumes of EM data and ease the burden of manual proofreading and annotation. Based on (Kaynig et al., 2015), we updated every stage of the software pipeline to provide better throughput performance and higher quality segmentation results. We used state of the art deep learning techniques to generate improved membrane probability maps, and Gala (Nunez-Iglesias et al., 2014) was used to agglomerate 2D segments into 3D objects. We applied the RhoanaNet pipeline to four densely annotated EM datasets, two from mouse cortex, one from cerebellum and one from mouse lateral geniculate nucleus (LGN). All training and test data is made available for benchmark comparisons. The best segmentation results obtained gave V Info scores of 0.9054 and 09182 for the cortex datasets, 0.9438 for F-score LGN, and 0.9150 for Cerebellum. The RhoanaNet pipeline is open source software. All source code, training data, test data, and annotations for all four benchmark datasets are available at www.rhoana.org